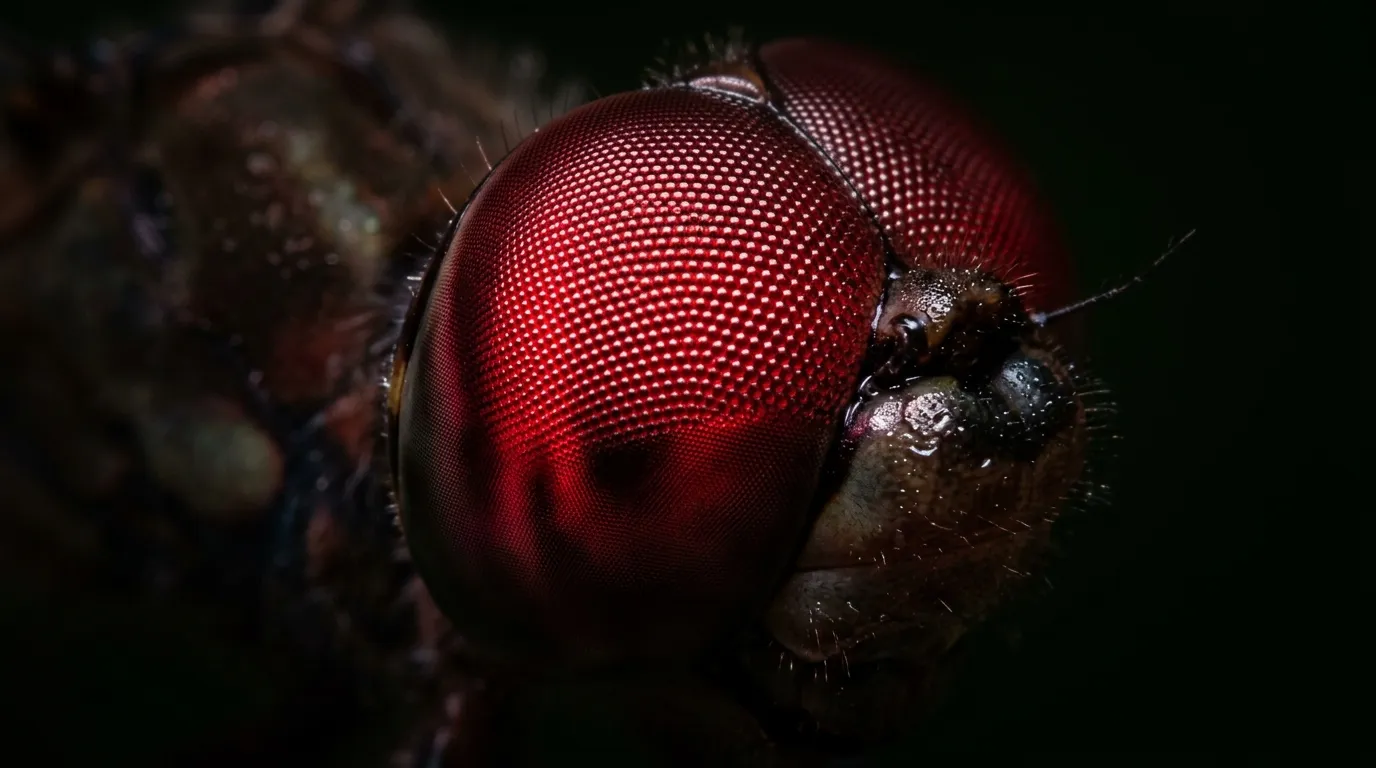
The dragonfly you watched last summer, hovering over a pond in hard afternoon light, was almost certainly seeing a color you cannot see. A team at Osaka Metropolitan University has just published the first direct demonstration that a family of common dragonflies uses a visual pigment tuned to roughly 720 nanometers, a wavelength so deep into the red that it is effectively at the mouth of the near-infrared range. Humans lose the ability to see anything at about 700 nanometers. The dragonfly, cruising above the cattails, is looking at a channel of the spectrum our eyes simply do not report.
What makes the finding unsettling, in a pleasing way, is not that the dragonfly sees further than we do. It is how it sees further. The paper, published this month in Cellular and Molecular Life Sciences by graduate student Ryu Sato and professors Mitsumasa Koyanagi and Akihisa Terakita, shows that the dragonfly's red opsin achieves its long-wavelength sensitivity through a molecular trick that is chemically identical to the one used by human red opsin. Two animal lineages that last shared a common ancestor more than 500 million years ago, somewhere in the pre-Cambrian murk, independently arrived at the same amino-acid configuration. The same engineering solution. The same protein fix. Twice.
Parallel evolution is one of the most quietly spectacular ideas in biology, and when it happens at the level of individual atoms in a protein, it is telling us something about the physics of the problem rather than the history of any particular animal.
What an Opsin Actually Is
To see why the Osaka finding matters, it helps to understand what the machine is. An opsin is a protein threaded through the membrane of a photoreceptor cell. It holds a small molecule called retinal, a vitamin-A derivative, in a snug pocket. When a photon hits the retinal, the molecule bends, and the opsin wrapped around it changes shape in response. That shape change triggers an electrical signal down the cell, which the brain eventually interprets as "I saw something."
The color of light an opsin is tuned to depends on the exact chemical environment around the retinal pocket. Different amino acids sitting near the retinal tug at its electron structure in subtly different ways. Pull harder, and the opsin absorbs shorter-wavelength light (blue). Pull gentler, and it absorbs longer wavelengths (red). Across all the animals that have been studied, opsin evolution is largely a story of tweaking the neighbors of the retinal to shift the absorption peak up or down the spectrum.
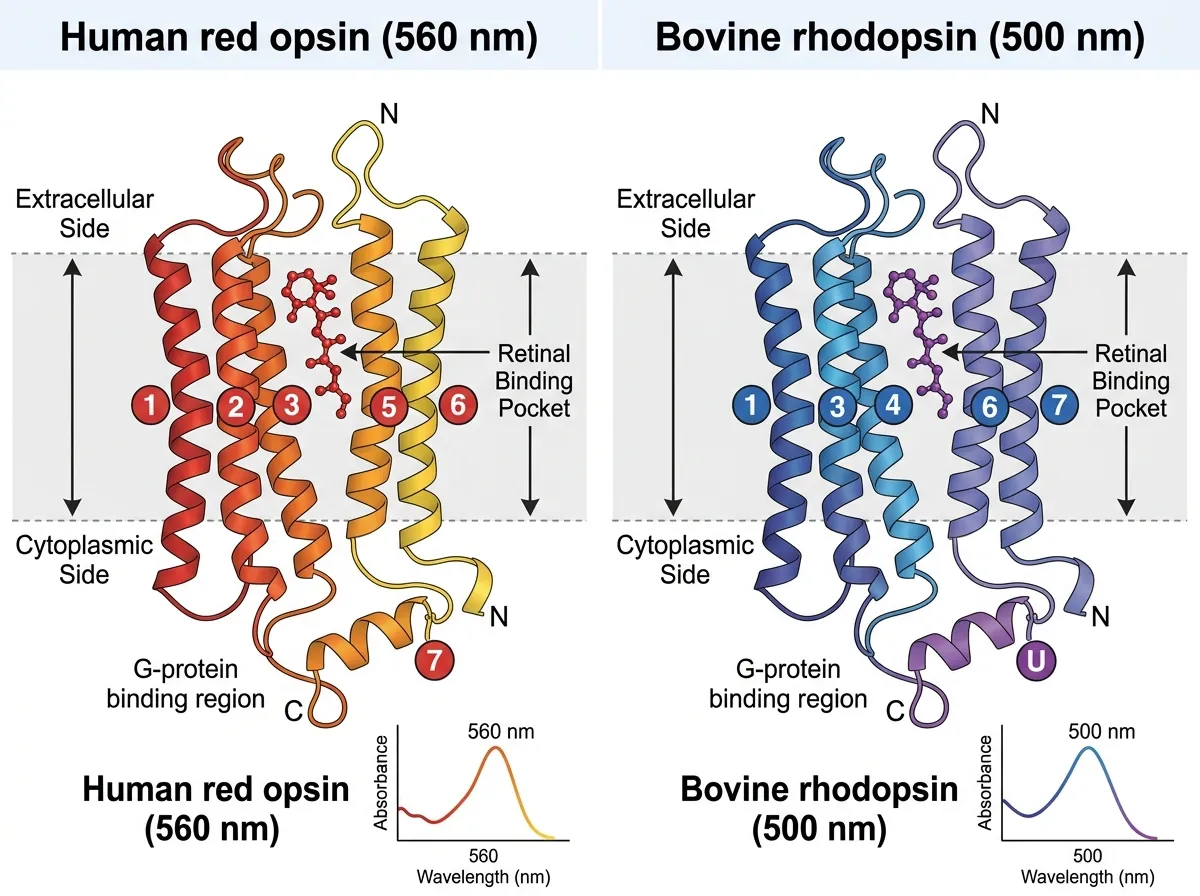
Humans have three color opsins: a short-wavelength one that peaks around 420 nanometers (blue), a medium one at about 530 nanometers (green), and a long one, the red opsin, peaking at 560 nanometers. Our "red" is not actually red. It is yellow-green. Red light only registers because the long opsin's sensitivity tail reaches into those wavelengths, barely.
Dragonfly vision, by contrast, is famously over-engineered. Different species carry up to 33 distinct opsin types, depending on the study you cite, which is more than any other animal known. Some of those are for ultraviolet. Some are for polarization. And at least one, the Osaka team has now confirmed, is for a red so deep it is essentially near-infrared.
The 720-Nanometer Question
When Sato and his colleagues began looking at dragonfly red opsins, they noticed something anomalous in one subfamily, the Gomphidae. The electrophysiological data from earlier field studies suggested Gomphidae dragonflies responded to wavelengths well beyond where any other insect red opsin had been documented. That kind of result is easy to hand-wave away as measurement artifact. The team wanted to know whether the molecular story matched.
They cloned the opsin genes, expressed the proteins in cultured cells, reconstituted them with retinal, and measured what wavelengths the purified pigments absorbed. One Gomphidae opsin peaked at 720 nanometers. For comparison, the reddest human opsin tops out at 560. The deepest red most people can perceive, above which the world goes dark, lies near 700.
Then they asked the key mechanistic question: how does the protein pull this off? Long-wavelength tuning in vertebrate red opsins had been traced, over decades, to a handful of specific amino acid substitutions. The most consequential of these is a hydroxyl-group-bearing residue that sits near the retinal pocket and stabilizes a positive charge on the retinal's Schiff base. The presence or absence of that hydroxyl, at that position, is the single largest determinant of whether a vertebrate opsin sees green or red.
The Osaka team sequenced the Gomphidae opsin, found the equivalent residue, and saw the same hydroxyl-bearing chemistry. They mutated it out. The absorption peak dropped toward shorter wavelengths, exactly as vertebrate opsins do when you perform the same mutation. The trick, at the level of individual atoms, was the same trick mammals use.
Parallel Evolution and the Grammar of Proteins
Evolutionary biologists split the duplication of form into three loose categories. Convergent evolution is when unrelated lineages arrive at similar outcomes from different starting points, like bat wings and bird wings: both are wings, but the underlying bone structure and developmental path differ. Parallel evolution is tighter. It is when lineages start from similar genetic raw material and arrive at the same solution through the same route. And molecular parallelism, the tightest case, is when the same amino acid changes occur at the same positions in homologous proteins.
The dragonfly-mammal red opsin story is molecular parallelism. It is the kind of result that makes theoretical biologists sit up, because it implies that the space of possible solutions to "how do I tune a protein to long-wavelength light" is not wide. For any opsin that needs to see red, one particular residue in one particular position is the cheapest lever. Evolution, being a cheapskate, finds that lever repeatedly.
Molecular parallelism has been documented in a handful of other systems. The same mutation that confers tetrodotoxin resistance in snakes preying on newts has shown up independently in other toxin-resistant species. The same lactase persistence variants appear in unrelated pastoral human populations. Certain color-vision mutations in fish recapitulate mammalian color-vision changes at the level of individual residues. Each case is an argument that the protein's function constrains the evolutionary possibilities more tightly than the sheer size of sequence space would suggest.
The point is not that evolution is deterministic. It is that at the level of molecules under strong selection, the number of roads up the hill is small. Cross-disciplinary patterns like this also appear in the history of thinking about neural function: we recently covered how astrocytes, long considered mere support cells, turn out to participate in memory formation in ways that echo neuronal mechanisms, a different flavor of the same story about function channeling form.
Why a Dragonfly Needs to See Into the Near-Infrared
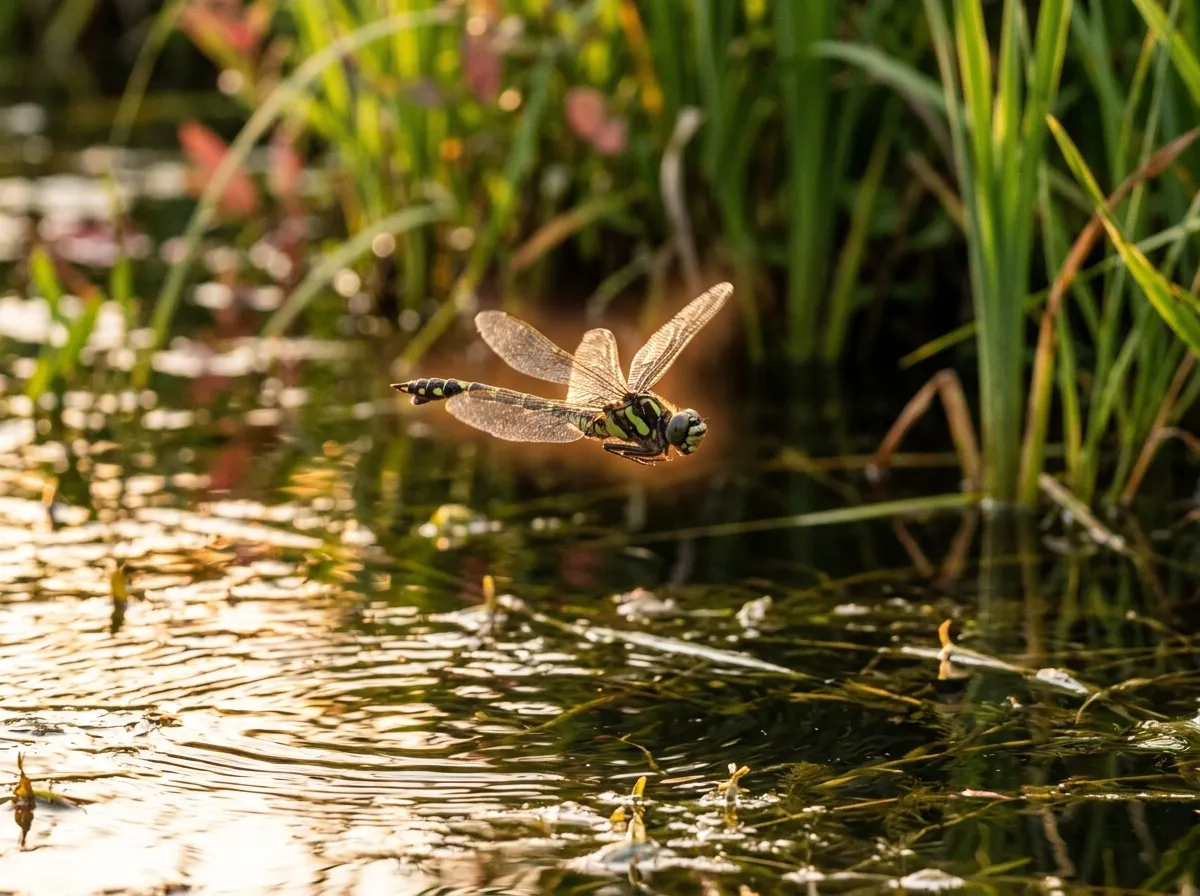
Knowing how an animal sees begs the question of why it bothers. Gomphidae dragonflies are shoreline hunters. They tend to perch on sunlit rocks and launch at prey flying above the water. The Osaka team, following up the molecular work with reflectance measurements, found that male and female Gomphidae bodies reflect substantially different amounts of light in the 700-to-720-nanometer range. A dragonfly with near-infrared sensitivity can tell sexes apart mid-flight in a fraction of a second, even when the scene is backlit by water.
There is also likely a thermal dimension. Long-wavelength light penetrates through haze and water vapor better than shorter wavelengths, which is why hunters and birdwatchers use near-infrared filters. For an insect chasing prey over a warm lake, extending the visual range slightly into the infrared may simply make the world less visually noisy. The evolutionary driver does not have to be dramatic. Modest advantages, applied millions of times over generations, are enough.
What the Osaka paper cannot tell us is how old the Gomphidae adaptation is. Dragonflies have been flying for roughly 325 million years, longer than trees have been a thing. Their opsin repertoire is ancient and tangled. Whether the 720-nanometer pigment is a recent invention of the family or a retained feature from a deeper ancestor is a question for comparative genomic work that has not yet been done.

The Optogenetics Payoff
The part of the Osaka story most likely to land in medical news is the engineering experiment the team ran after the biology. They took the Gomphidae opsin, mutated a single additional residue, and shifted its sensitivity even further toward longer wavelengths. The modified opsin produced measurable cellular responses when illuminated with near-infrared light. That is a useful tool.
Optogenetics is the technique that lets neuroscientists turn specific neurons on or off with light, by genetically inserting light-sensitive proteins into those cells. The technique has transformed basic brain research over the past fifteen years. Its main practical limitation is that most opsins currently in the optogenetics toolkit respond to blue or green light, which does not penetrate tissue well. Any attempt to activate neurons deep in the brain of a living animal, or deeper than a few millimeters into any biological tissue, runs into a hard absorption wall.
Near-infrared light does not hit that wall. It passes through centimeters of tissue with minimal scattering and absorption. An opsin tuned to near-infrared could, in principle, be activated through intact skull and skin. For therapeutic applications in conditions like Parkinson's disease, epilepsy, or chronic pain, that difference between "works through a thin layer of cells" and "works through a whole head" is enormous.
"This is one of the most red-sensitive visual pigments ever found," Terakita said in the press release accompanying the paper. Whether the Gomphidae opsin, engineered or not, becomes a standard piece of the neuroscience toolkit is a question that will take years to answer. What the paper does is open the possibility.
The Deeper Question
Every so often a biology paper arrives that is interesting in three distinct ways at once. The Osaka study is one of those. It is a molecular biology result, showing a specific protein doing something unusual. It is an evolutionary result, showing the same unusual thing being discovered twice. And it is a tool-development result, handing optogenetics researchers a new color of light to work with.
What unites the three, and what is probably the lasting contribution of the work, is the reminder that proteins have a grammar. The sentences evolution can write are not infinite. When you need to pull a retinal molecule's absorption spectrum out to 720 nanometers, there is a small vocabulary of amino-acid substitutions that will do the job, and every lineage that ends up with long-wavelength vision ends up reaching for the same words. A dragonfly hovering over a Japanese pond and a primate reaching for red fruit in an African forest turn out to be running the same chemical program.
The remaining puzzles are big. Why Gomphidae and not other dragonfly families? How many times has this same substitution been found and lost across invertebrates? What other near-infrared visual systems are out there, unsurveyed, in species we have not thought to ask? The paper is a beginning, not a conclusion.
But it is a beginning that tells us, once again, that the natural world has solutions we have not noticed, built out of letters we already know.
Sources
- Sato R, Terakita A, Koyanagi M. "Dragonfly red opsins share a common tuning mechanism with mammalian red opsins and further enhancement of near-infrared sensitivity." *Cellular and Molecular Life Sciences*, 2026.
- ScienceDaily, "Dragonflies can see a color humans can't and it could change medicine"
- EurekAlert, "Seeing red: Dragonflies and humans share the same red-light detection mechanism"
- Neuroscience News, "The Shared Secret of Mammal and Dragonfly Color Vision"
- Sci.News coverage of the Osaka opsin finding

